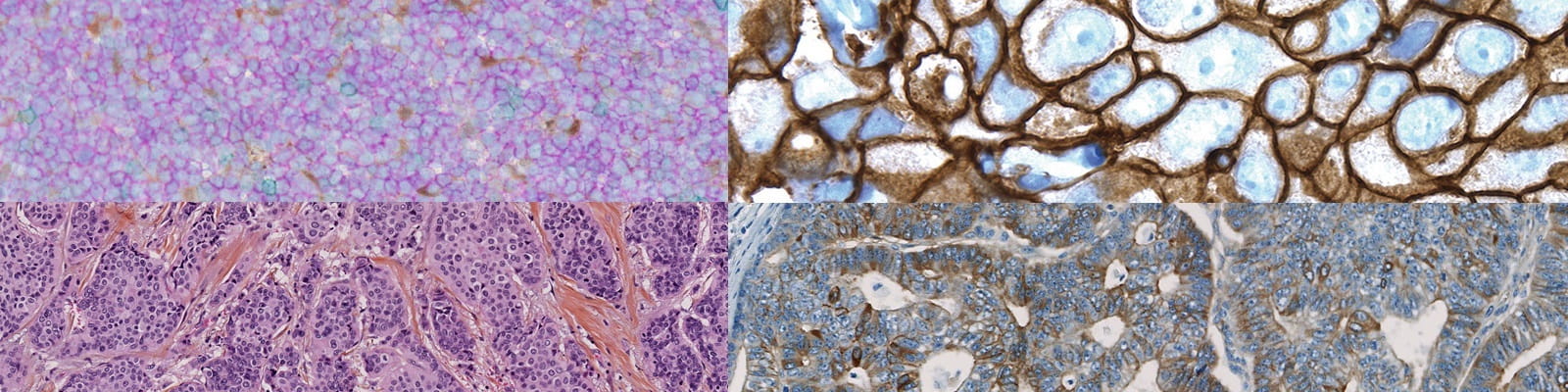
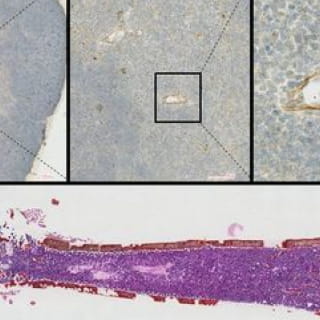
Downregulation of stromal syntenin sustains AML development.
Leblanc R, Ghossoub R, Goubard A, Castellano R, Fares J, Camoin L, Audebert S, Balzano M, Bou-Tayeh B, Fauriat C, Vey N, Garciaz S, Borg JP, Collette Y, Aurrand-Lions M, David G, Zimmermann P